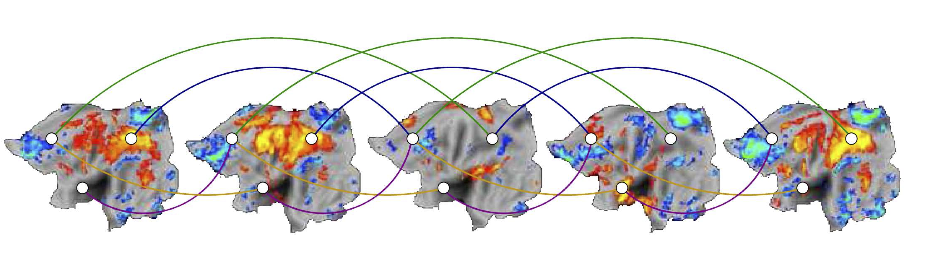
Connectome Graph based Source Modeling
Inferring patterns of cortical activity from EEG measurements taken on the surface of the scalp is a challenging inverse problem. The fundamental challenge is that source currents are not observed directly, and that number of degrees of freedom of possible cortical activity is much higher than the number of electrodes that may be placed on the head. A common approach to dealing with this issue is to employ some prior model for the cortical sources, such as the belief that "good" cortical activity solutions will be smooth, or have small norm.
We seek to construct such cortical source prior models that are consistent with the connectivity of the brain. Recent advances in MRI imaging modalities (notably DTI and DSI) have enabled the measurement of directionality of white matter fiber tracts in the brain. Combined with fiber tractography methodologies, these technologies allow measurement of the density of connections between brain regions, which may represented as a weighted graph. Our key idea is that brain activity is likely to be similar in highly connected regions. In the simplest case, we construct a penalty functional for cortical activity by summing the squared differences across the edges of the connectome, thus penalizing patterns of cortical activation inconsistent with measured anatomical connectivity. We are also exploring spatiotemporal extensions that allow modeling of signal delays, which we expect to be significant at high temporal frequencies relevant for EEG.